Why Are Bacteria So Simple?
As far as we can tell, bacteria were the first lifeforms on Earth. Which means they’ve had a full four billion years to make something of themselves. And yet, despite their long evolutionary history, they mostly still look like this:
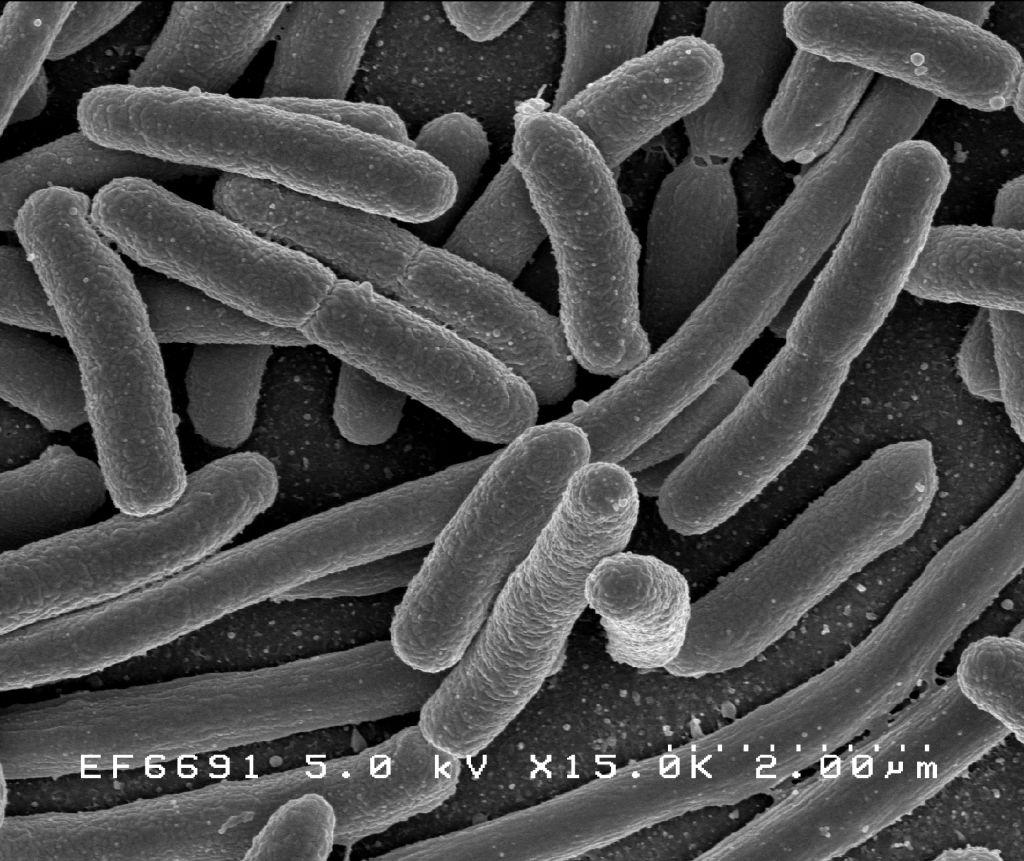
Bacteria belong to one major class of cells—prokaryotes.[1] The other major class of cells, eukaryotes, arrived about one billion years after bacteria. But despite their late start, they are vastly more complex.
Prokaryotes mostly only contain DNA, and DNA translation machinery. Eukaryotes, on the other hand, contain a huge variety of internal organelles that run all kinds of specialized processes—lysosomes digest, vesicles transport, cytoskeletons offer structural support, etc.
Not only that, but all multicellular life is eukaryotic.[2] Every complex organism evolution has produced—eukaryotic. Trees, humans, worms, giant squid, dogs, insects—eukaryotic. Somehow, eukaryotes managed to blossom into all of these complex forms, while bacteria steadfastly remained single-celled, simple, and small. Why?
The short answer is that prokaryotes have vastly less DNA than eukaryotes—four to five orders of magnitude less, on average—and hence can’t do nearly as much stuff.[3] The long answer is the rest of this post, which investigates two related questions: first, why are eukaryotic genomes so long? And second, how exactly does more DNA allow for more complexity?
Why Are Eukaryotic Genomes So Long?
Scalable Energy Production
Using DNA—replicating, transcribing, and translating it into proteins—isn’t free. Cells need energy (such as ATP) to power these reactions and, all else equal, longer genomes will require more of it.[4]
Both prokaryotes and eukaryotes pay similar energetic costs to maintain genes. The difference is that eukaryotes have way more energy and hence can afford to have longer genomes. But why this disparity?
Prokaryotes generate ATP along their cell membrane. Which means that as they increase in size, their surface area—and hence their energy production—will scale sublinearly with their volume. So a prokaryote that doubles in size, for example, will only end up producing half as much ATP per unit volume. Because prokaryotes become less metabolically efficient as they get bigger, most are quite small—six orders of magnitude smaller than eukaryotes, on average.[5]
There are some exceptions. For instance, individual bacteria in the species Thiomargarita can reach up to one centimeter in size, visible to the naked eye! But its cell structure suggests the exception proves the rule—80% of its volume is a vacuole,[6] essentially empty space. So in effect, evolution expanded its surface area without concomitantly expanding its functional volume—a neat trick!
But how do eukaryotes avoid this surface area constraint? Well, eukaryotes generate energy using mitochondria, which are inside the cells. As a result, their number of mitochondria—and hence their energy production—scales with their volume. This allows them to afford both larger cell sizes than prokaryotes, and also longer genomes.[7]
Tolerance for Junk
But bioenergetic constraints aren’t the whole story. Even leaving aside the direct energy costs, prokaryotes face way more selection pressure toward having short genomes.
Empirically, bacteria are very quick to rid themselves of genes once they’re no longer useful. For example, if you insert DNA into a bacteria that affords antibiotic resistance, it will keep those genes as long as antibiotics are around. But once you remove the antibiotics, it will jettison that DNA within a few hours.[8]
Eukaryotic DNA, on the other hand, is much more weakly selected against. While bacteria are sensitive to additions of DNA fewer than ten base pairs in length, eukaryotes will keep additions of over ten thousand around indefinitely, even if they’re useless.[9]
But why is selection so much weaker among eukaryotes? The main reason is that they have very small population sizes relative to bacteria, and the smaller the population size, the more the species’ genome will be determined by chance. This sentence requires a bit of unpacking.
What does it mean for a genome to be determined “by chance”? There are, generally speaking, two ways by which new genes can spread throughout the population. They can be actively selected for, or they can propagate purely by random events (also referred to as genetic drift).
The likelihood that a gene spreads through the population by chance alone is inversely related to the size of the population. After all, the gene faces the same probability of propagation at each reproduction event, so the more individuals, the more unlikely it is to reach all of them. Conversely, the smaller the population size, the more likely genes are to spread by chance.[10]
Of course, selection will still promote genes with high fitness and cull those of low fitness. But in the vicinity of neutral fitness, the effects of chance begin to dominate a gene’s fate. And as population sizes decrease, this vicinity grows, and the fate of more genes will be determined by chance. Put differently, as population sizes decrease, selection becomes a weaker force.
Prokaryotes have vastly larger population sizes than eukaryotes,[11] which means that their genomes are under stronger selection—any non-immediately useful gene is quickly discarded. Eukaryotic organisms, on the other hand, often have such small population sizes that large portions of their genomes evolve almost as if natural selection were entirely absent. This means that even mildly deleterious stretches of “junk” DNA will tend to stick around and accumulate.
But why, one might wonder, do eukaryotes have such small population sizes? The main reason seems to be that eukaryotic organisms are much bigger, and bigger animals tend to have smaller population sizes. In general, if an organism becomes twice as large, the overall population of such organisms will halve.[12] I don’t know why this is true, although I suspect it has to do with food supply: as organisms get bigger, they each need more energy from their environment and there is only so much to go around, so the total number of individuals shrinks.
How Do Longer Genomes Enable More Complexity?
So, eukaryotic genomes will tend to keep superfluous DNA around. But how does that DNA actually get there in the first place? After all, if mutations are just as likely to be deletions as they are insertions, then the net effect on genome length should be zero.
Unfortunately for everyone, one of the main ways DNA lengthens is that genetic parasites called transposons[13] copy and paste themselves throughout the genome. Indeed, at least 45% of the human genome is the result of these fuckers.[14] The other major source of genome expansion is thankfully less depressing—just some accidental duplication, typically of a single gene, but occasionally of an entire chromosome!
As we saw above, prokaryotes don’t stand for this kind of fuckery—any bit of DNA which isn’t immediately useful is typically discarded. Thus, parasitic DNA and accidental duplications are very unlikely to accumulate.
Eukaryotes, on the other hand, will keep loads of this random, completely unhelpful DNA around. So while nearly all DNA in prokaryotes is protein-coding, in eukaryotes—and especially in highly complex organisms like humans—sometimes as little as 1% of the genome codes for any protein.
Eukaryotic Monopolies
At this point you might be wondering, wait, wasn’t more DNA supposed to entail more complexity? It seems like eukaryotes just got the shit end of the stick—a ton of parasitic junk DNA that is at best useless and at worst mildly deleterious, but just barely not deleterious enough for selection to notice. This is supposed to be… good?
Yes and no. While it’s true that a huge fraction of eukaryotic DNA is almost certainly just “junk,” the slack on their genome size also creates more opportunity for innovation. It is precisely because eukaryotic DNA is long and under little selection pressure that it has the chance to evolve useful secondary adaptive changes later on.
Peter Thiel argues that monopolies often actually spur innovation. While companies caught in cut-throat markets need immediate returns, monopolies have the financial freedom to pursue basic research that might pay off in the future. Eukaryotes, in this borrowed metaphor, are more monopolistic, innovating over longer time horizons, while bacteria are limited to lives of myopia, only creating products which can pay off quickly.
But how is it that these initially-useless mutations come to pay off later? And what kinds of innovations did eukaryotic genomes “invent”?
Duplication and Divergence
At a high level, the main difference between prokaryotic and eukaryotic DNA is that the latter has way more “software.” What is “software” in a genome? As a very crude analogy: genes which code directly for proteins can be thought of as hardware, and genes which regulate what those protein-coding genes do can be thought of as software.[15]
One way that eukaryotic genomes can acquire this software is through a process called “duplication and divergence.” The basic idea is that sometimes a regulatory gene is accidentally duplicated. In bacterial genomes, these duplications would almost certainly be quickly deleted—it’s just extra clutter that costs precious energy to maintain for no additional benefit. But eukaryotic DNA is fine with all of these mildly deleterious additions! So these duplications can stick around, being redundant for a while.
But slowly, this might start to change. Let’s say, for example, that there is a protein-coding gene which codes for a stress hormone, and a regulatory gene which can turn that gene “on or off,” i.e., control when that hormone is created. Originally, this regulatory gene only activates the hormone in liver cells in the presence of toxins. But once the regulator has been duplicated, the second copy may begin to mutate away from its original function. Perhaps the mutation now causes it to activate the same hormone in a different cell type—in skin cells when they’ve been bruised, for example. In this way, duplicated regulators may occasionally “diverge,” i.e., take on new functional roles.[16]
Duplication and divergence is one mechanism whereby software proliferates in eukaryotic genomes. But eukaryotes also have better software. In particular, eukaryotic genomes have more modularity and higher-level abstractions. Prokaryotes, on the other hand, cannot get past the duplication step, and so these routes to complexity are unavailable to them.
Modularity
To the extent that prokaryotic genomes do have regulatory elements, these often control “operons,” sets of genes which are all turned on or off at the same time. In other words, none of the genes in the set can act independently of each other.
Operons typically achieve a single function. For example, E. coli prefers glucose as its energy source, but when glucose is in short supply it will switch to using lactose. This is controlled by the lactose operon: when regulatory elements sense that glucose is absent and lactose is present,[17] they will trigger the expression of a set of genes which jointly work to create the proteins necessary to digest lactose.[18]
Eukaryotic genomes are almost entirely devoid of operons. Instead, their genes are modular, in the sense that any single gene is typically used in many different operations, rather than being part of one functional unit. This sort of modularity enables the underlying “hardware” to be used much more flexibly.
And this flexibility is a large part of what allows for multicellularity. Different cell types (e.g. neurons, liver cells) within the same organism are defined not by different genomes, which are the same in all cells, but by different patterns of gene expression. These patterns can be staggeringly complex, but at a very basic level they are just regulatory genes processing information (e.g., sensing that lactose is present) and controlling the timing and amount of protein-coding genes (or other regulatory genes!) in response.
Because eukaryotic genes are so modular, the space of possible genetic patterns is massive, and hence the space of possible phenotypic structures is massive, too. When you combine this with the fact that prokaryotic genomes contain far fewer protein-coding genes (and even fewer regulatory ones), the difference in the amount of phenotypic possibilities is truly staggering.
At its core, multicellularity results from different patterns of gene expression. Technically, this is possible with operons: just mix and match them in different ways! But this tool is far more crude. Instead of being able to adjust patterns at the gene level (and hence the individual protein level), patterns which utilize operons are restricted to operating at the circuit level (e.g., digest lactose), and hence the space of possible prokaryotic phenotypes is dramatically reduced.
How might modularity emerge from the duplicate and diverge mechanism? It’s the same idea as the liver and skin example explained above. A regulator controls a protein-coding gene in one context (the liver) and a duplicated regulator eventually mutates to the point that it controls it in a different context (the skin). Now the gene is being used independently in two different patterns: it’s modular!
Abstractions
In addition to modularity, eukaryotic genomes also operate at higher levels of abstraction. For instance, there are the so-called “master genes” which can induce macroscopic body structure. One example is the Pax6 regulatory gene. Forcing this single gene to be expressed where it normally wouldn’t be—e.g., in the legs or abdomens of flies—causes an entire eye to form there.[19]
This means evolution does not have to work from scratch every time to create new body plans. Instead, small tweaks can be made to high-level variables (like master genes) to create novel macroscopic structure, such as adding additional legs, or moving the position of eyes. Indeed, the diversity of body plans among animals seems to stem primarily from using the same underlying regulatory genes in different patterns.[20]
How might these higher-level abstractions arise in practice? I’m not sure, but here’s a (speculative) sketch of my guess:
Say you have two genes, each of which makes a particular protein. The first gene creates a hormone that causes food-seeking behavior; the second an enzyme that breaks down glucose. And suppose these two functions happen to be synergistic, in the sense that whenever the cell is stressed it should both stop seeking food and stop using glucose.
At first these two genes were regulated separately. But suppose one of the regulators gets duplicated, and then eventually mutates, such that it gains the ability to control both genes at once. Now, many sub-functions are regulated by a single gene, enabling it to represent information at a higher-level of abstraction, e.g., about the overall stress level of the organism.
At this point the process can repeat, at increasingly higher levels of abstraction. Eventually long chains of these regulators regulating other regulators might form, enabling hierarchies of subroutines that master genes can control—affording single genes the ability to create entire macroscopic structures, like eyes.
Prokaryotes don’t have these high-level genes for the obvious reason—there is nothing high-level to control! But this answer passes the buck. There is nothing high-level for them to control because they don’t have the tools to build complex structures in the first place. Bacteria didn’t really get modularity or high-level abstractions—bacteria got hardly anything—because they can’t keep useless genes around long enough for secondary adaptive changes to emerge later on.
So, Why Are Bacteria So Simple?
We can now finally say why bacteria are so embarrassingly simple. There are two pressures keeping their genomes short—lack of energy and strong selection—and this shortness does indeed limit their ability to build complex structures. Without the slack to explore a wider range of regulatory possibilities, their software is stunted—indeed, almost non-existent. Couple this with the fact that they have far fewer coding genes to begin with, and we have our result: four billion years of potential with only some tiny boring blobs to show for it.
And while eukaryotic genomes might, at first glance, seem undesirable—bloated with junk which is at best redundant and at worst parasitic—with that bloat comes length, and length can do all kinds of wonders over evolutionary time. Not only do longer genomes give eukaryotes the chance to accumulate more protein-coding genes, but they also enable software upgrades like modularity and higher-level abstractions. And from these, complexity follows: as software becomes more hierarchical, with more flexibility over the underlying hardware, innovations such as multicellularity and the staggering diversity of body plans become possible.
It’s kind of insane how complex eukaryotic organisms got, considering that they started out as tiny little blobs stuck in the muck, just like bacteria. To be sure, they’ve been around a long time, but as we’ve seen time isn’t all that matters—bacteria had a one billion year head start and yet stuck in the muck they remain. Biological complexity, it seems, is about more than just time and chance: it is about exploration, the ability to try, but more importantly the ability to fail, so that when something useful does finally come around, it can be seized upon. So that eukaryotes can slowly claw their way out of the muck, can complexify…
… while their bacterial cousins continue to press “exploit” for eternity.
Thank you to Adam Scholl for invaluable writing feedback and for countless fun and thoughtful conversations (I promise I’ll stop talking about bacteria now :p), to Alexander Gietelink Oldenziel for suggesting really useful books and papers on the topic, for helping think through some of the trickier population genetics claims, and for feedback on earlier drafts, and thank you to Siddharth Hiregowdara for feedback on a previous draft.
- ^
The technical distinction is that eukaryotes have a nucleus—a small compartment which holds the DNA. Prokaryotes don’t, their DNA floats freely around inside the cell.
- ^
There actually are some cases of prokaryotic multicellularity. For instance, bacteria will sometimes aggregate together in what are called biofilms. Biofilms can exhibit cooperative behavior—e.g., cells on the inside of the film will send out signals of starvation to the outside, causing the exterior cells to halt activity and wait until the interior ones are fed. There are also cases of cell specialization among prokaryotes. For instance, in one bacterial species (Nostoc) there is a cell type which has specialized to metabolize nitrogen. But aside from the occasional multicellular blob, bacteria mostly remain single-celled and simple creatures. They never saw the explosion of intelligence, complexity, and diversity of body plans that eukaryotes did.
- ^
There are many estimates of this, but e.g.: mitochondria “enabled a roughly 200,000-fold rise in genome size compared with bacteria” (Lane and Martin).
- ^
These costs are not trivial. As a rough estimate: an average E. coli gene costs around one ten thousandth of the cell’s total energy budget to maintain. Given that there are around 4,000 genes in E. coli, the entire genome adds up to about a tenth of its total energy budget (Lynch and Marinov).
- ^
This is only the disparity between prokaryotes and single-celled eukaryotes; a similar disparity exists between single-celled eukaryotes (such as yeast) and multi-celled ones. (See for example The Origins of Genome Architecture, page 83, under the section “The Three Genomic Perils of Evolving Large Body Sizes”).
- ^
“Cells showed a large central vacuole which accounted for 73.2 ± 7.5 % of total volume” (Volland et al.).
- ^
The full extent of this argument is made in Nick Lane’s book Power, Sex, and Suicide: Mitochondria and the Meaning of Life. See in particular, the chapter “The Foundations of Complexity.”
- ^
See Nick Lane’s book Power, Sex, and Suicide, page 118, section “Balancing gene loss and gain in bacteria.”
- ^
See the section on “Gene Structural Costs” in Lynch and Marinov.
- ^
In particular, the chance that a gene spreads throughout the population by chance alone is equal to 1/N, where N is the effective population size (explained in next footnote). To see why, consider that each new allele introduced to the population has the same chance of going to fixation (ignoring selective forces). Since they all have the same chance, and since there are N copies of different alleles currently in the population, the chance that this new, unique allele spreads to everyone by chance alone is 1/N. Thanks to Alexander Gietelink Oldenziel for this argument.
- ^
Prokaryotes typically have effective population sizes of 10^9. The effective population size takes into account factors like sex ratio, geographic distribution, etc., and so is often much smaller than the total population size. Eukaryotic organisms have effective population sizes ranging from 10^4 for invertebrates like us to 10^7 for single cells like yeast (Lynch and Marinov). To be clear, these variations in population size have real, substantial effects on the force of selection; refer to the paper for a great in depth analysis of this.
- ^
See Michael Lynch’s book The Origins of Genome Architecture, page 84, under the section “Smaller population size.”
- ^
These go by many other names, e.g.: jumping genes, selfish genes, and mobile genetic elements.
- ^
See Sean Eddy.
- ^
This gets quite complicated, but you are presumably in the footnote section for more complicated answers, so I will gander to tell you. For one, the hardware/software distinction is not totally apt. After all, the protein-coding genes are “codes” for the protein, not the protein itself. Ah well, for a very crude analogy it probably holds up.
Second, what does it mean for a regulatory gene to “control” what a protein-coding gene does? Well, every protein-gene has what is called a promoter sequence that sits directly above the gene. This sequence does not code for proteins, it is just a stretch of DNA which is particularly attractive to the molecules which start the “coding for protein” process. So, whenever this is exposed, the protein-coding gene is more likely to produce proteins. And conversely, whenever it is blocked, the gene can’t produce proteins. Now genes can be turned “on or off” by blocking or unblocking the promoter.
Regulatory genes do just this: they can, e.g., create proteins which serve the functional purpose of blocking the promoter. Or they can be non-coding, e.g., stretches of DNA like the promoter that bind well to molecules which affect genetic expression. This is somewhat unfortunate, since there is not a clean distinction between “protein-coding” and “regulatory,” as some regulatory genes do make proteins! Gah. In general, though, it’s my impression that regulatory elements tend to be of the non-coding variety.
- ^
See Michael Lynch’s book The Origins of Genome Architecture, page 294, under the section “The passive emergence of modularity.”
- ^
Here, what it means for a “regulatory element to sense,” is that a protein created by a regulatory gene can bind to lactose, and when it does, that protein changes shape such that it detaches from the promoter sequence (see footnote 14 for more details) and the operon can be expressed.
- ^
See Essential Cell Biology (fifth edition), page 294, under “How Transcription is Regulated” for more details on this example and operons in general.
- ^
See From DNA to Diversity, page 29, section on “Field-specific selector genes” for more information and pictures of this process.
- ^
See From DNA to Diversity for many such examples, in particular the section “Sharing of the genetic toolkit among animals.”
So does this mean that when bacteria evolve antibiotic resistance in a new location (i.e. new hospital, new farm, etc), they have to do it de novo?
Good question! I don’t know, but I think that they don’t necessarily need to. Something I didn’t get into in the post but which is pretty important for understanding bacterial genomes is that they do horizontal gene transfer, which basically means that they trade genes between individuals rather than exclusively between parents and offspring.
From what I understand, this means that although on average the bacteria shed the unhelpful DNA if given the opportunity, so long as a few individuals within the population still have the gene, it can get rapidly reacquired when needed. I don’t know exactly how the math works out, but I’d guess that in big enough populations, if antibiotic encounters are somewhat common, then probably they don’t need to do it de novo each time?
This also means bacterial genomes are much more distributed than eukaryotic ones. So long as any individual bacteria has some gene, it’s “as if” the whole species has it. Which means their genomes are, in a sense, actually longer than they might naively seem. Being distributed has advantages: no single genome needs to be very long, yet the population can hold onto useful stuff. But it also has disadvantages: any adaptation that relies on genes being close together in a single genome is unlikely to develop (which includes e.g. all of the regulatory hierarchy stuff mentioned in the post). So I do still expect that the pressure towards short genomes meaningfully stunts bacterial complexity.
Good question! There’s a literature on the epidemiology of drug-resistant bacteria, and they do manage to get transmitted between hospitals and across international borders in the bodies of hosts.
So unfortunately no, we can breed antibiotic resistant bacteria in one place and spread them around all over.
wait so is a viable strat re: antibiotic resistant infections to just go a while without any antibiotics, then restart? granted, the patient needs to survive, and ideally not spread the infection to others, but still. Is this a documented action suite?
I don’t know for sure. It seems important that most antibiotic resistant infections start in a patient who is not on antibiotics, suggesting there’s a clinically meaningful time delay before the bacterial population loses antibiotic resistance in the absence of selection pressure.
Hospitals have tried approaches of either using distinct antibiotics on different patients, or rigidly using one antibiotic at a time on all patients on a rotation, in order to avoid antibiotic resistance, and I know the efficacy of these approaches does get studied.
I think with antibiotic resistance, we see the doctors trying one drug after another on the patient until they see a response. For each drug, the bacteria in the patient will have gone at least the length of time since the patient’s infection started since last being exposed to it. Possibly a patient who responds to no drugs but who manages to be kept alive despite the infection could then find success if the doctors tried one of the old drugs again. My guess is this is just a rare circumstance. In most cases the patient will have recovered or died.
Think of it as exponential decay. If the half life of a completely non-functional gene is only 2 hours, then after a week only one in 10^25 bacteria have it—i.e. none. Trace amounts of functionality (like trace amounts of antibiotics) will slightly extend the half life—but this causes outsized effects at the tails. If the half life of an antibiotic resistance gene when there’s trace antibiotics gets extended to 3 hours, then after a week one in 10^17 bacteria still have it—i.e. still probably none, but maybe a few that can carry the gene from farm to farm or hospital to hospital.
One other mechanism that would lead to the persistence of e.g. antibiotic resistance would be when the mutation that confers the resistance is not costly (e.g. a mutation which changes the shape of a protein targeted by an antibiotic to a different shape that, while equally functional, is not disrupted by the antibiotic). Note that I don’t actually know whether this mechanism is common in practice.
So happy to see this post appear! 🔥
The story about operons and the high interconnectedness of prokaryote genomes makes me wonder: bacteria kick out the antibiotic-coding gene after a few hours… but how does it know which gene to kick out?
Does it have a way to tell which genes are more ‘alien’ than others? (Or are we only talking about plasmids here?) I’ve heard it’s hard to genomic manipulate some genomes because the cells keep kicking out new genes
One could speculate there is some sort of mechanism, perhaps epi-genetic, that is able to tell which genes are more alien / new than others somehow?
I’d love to hear your thoughts
Bacteria have systems such as CRISPR that are specialized in detecting exogenous DNA such as from a potential viral infection.
They also have plasmids that are relatively self-contained genetic packets, which are commonly the site of mutations conferring resistance, and which are often exchanged in the bacterial equivalent of sex.
However, to the best of my knowledge, there’s no specific mechanism for picking out resistance genes from others, beyond simple evolutionary pressures.
The genome is so small and compact that any gene that isn’t ‘pulling its weight’ so to speak will likely be eradicated as it no longer confers a survival advantage, such as when the bacteria find themselves in an environment without antibiotics.
Not to mention that some genes are costly beyond the energy requirements of simply adding more codons, some mechanisms of resistance cause bacteria to build more efflux pumps to chuck out antibiotics, or to use alternate versions of important proteins that aren’t affected by them. Those variants might be strictly worse than the normal susceptible version when antibiotics are absent, and efflux pumps are quite energy intensive.
There’s no real foresight involved, if something isn’t being actively used for a fitness advantage, it’ll end up mercilessly jettisoned .
Thanks!!
Yeah I think it’s a great question and I don’t know that I have a great answer. Plasmids (small rings of DNA that float around separately) are part of the story. My understanding here is pretty sketchy, but I think plasmids are way more likely to be deleted than the chromosomal DNA, and for some reason antibiotic resistant genes tend to be in plasmids (perhaps because they are shared so frequently through horizontal gene transfer)? So the “delete within a few hours” bit is probably overstating the average case of DNA deletion in bacteria. I would be surprised if it “knew” about the function of the gene, although I agree it seems possible that some epigenetic mechanism could explain it. I don’t know of any, though!
I assume this is how we make AGIs controllable right. Give them tight constraints on their network size and weights and execution depth and short term memory. Force them to optimize for short term exploiting.
This prevents most hostile plans as something like “take over the planet and then make more paperclips” has a period of time where less paperclips are being made than the immediate term “order more wire and paperclip bending robots”.
Alas, the commands to open a cloud computing account and allocate a million VMs are very low-energy. AI can scale in ways that living things cannot.
One thing worth mentioning is that proto-eukaryotic cells originally got the energy to focus on larger scale things by capturing other prokaryotic cells. Mitochondria and chloroplasts look a lot like simplified bacteria that specialise in producing ATP and glucose+oxygen, respectively, both of which give the cell more energy. They even have their own, specific DNA inside them.
To get really speculative, this is quite similar to how agriculture and the resulting centralisation+specialisation allowed for the explosion of civilization, where the last ~10k years have overwhelmingly more complex stuff than the preceding 100k years.
Your link to Lynch and Marinov is currently incorrect. However I also don’t understand whether what they say matches with your post:
Ah, thanks! Link fixed now.
Yes, welp, I considered getting into this whole debate in the post but it seemed like too much of an aside. Basically, Lynch is like, “when you control for cell size, the amount of energy per genome is not predictive of whether it’s a prokaryote or a eukaryote.” In other words, on his account, the main determinant of bioenergetic availability appears to be the size of the cell, rather than anything energetically special about eukaryotes, such as mitochondria.
There are some issues here. First, most of the large prokaryotes are outliers like Thiomargarita, in the sense that they have expanded their energy without expanding their functional volume. However, their genomes are still quite small, which means that their “energy/genome” will be large. Eukaryotic cells of the same size have way more energy and way longer genomes, making their “energy/genome” roughly equivalent to the large prokaryotes.
Second, Lynch’s story is that strong selection keeps bacterial genomes short. The main reason that bacteria have strong selection is because there are so many of them, and there are so many of them because they’re so small. But why are they so small? It seems like an obvious contender is Lane’s story about them being energy bottlenecked by their surface area. So, in my opinion, these two hypotheses are synergistic and my best guess is that they’re both part of the story.
You may find this paper interesting. Among other things, they find that total population metabolism (i.e., individual basal metabolic rate times the population size divided by the area occupied by that population) stays close to constant relative to body mass, when measured across all eukaryotes.
So eukaryotes have nuclei, but the thing that makes them special is the mitochondria. Do we see anything that has a nucleus but not mitochondria, or vice versa?
We know of at least one eukaryote species that lost its mitochondria. I don’t know about mitochondria but no nucleus. Erythrocytes don’t have either, so maybe (speculating here) there are some specialized animal cells that have mitochondria but no nuclei.
The details are good, but I reject the framing. Bacteria are simple because simple outcompetes complex. Eukaryotes fill a small niche. We care about complexity because we are in that niche. Phrased that way, it’s not surprising that only a single lineage fills that niche and that it took a billion years to try it.
If simple outcompetes complex, wouldn’t we expect to see more prokaryotic DNA in the biosphere? Whereas in fact we see 2-3x as much eukaryotic DNA, depending on how you count—hardly a small niche!
The LessWrong Review runs every year to select the posts that have most stood the test of time. This post is not yet eligible for review, but will be at the end of 2024. The top fifty or so posts are featured prominently on the site throughout the year.
Hopefully, the review is better than karma at judging enduring value. If we have accurate prediction markets on the review results, maybe we can have better incentives on LessWrong today. Will this post make the top fifty?
Nice post!
I feel like the real question to answer here isn’t “Why are bacteria so simple?” (because if they were more complex they wouldn’t really be bacteria anymore), but rather “Why do there seem to be those 2 classes of cells (eukariotes and prokariotes)?”. In particular, (1) why aren’t there more cells with intermediate size and complexity, and (2) why didn’t bacteria get outcompeted out of existence by their cousins which were able to form much more complex adaptations?
(Note: I know very little about biology. Don’t trust me just because I never heard of medium-sized and medium-complex cell types that don’t neatly fit into one of the clusters of prokariotes and eukariotes.)
Perhaps software/firmware instead of software/hardware?
That’s fascinating… do we understand the mechansim by which they correctly “determine” that this DNA is no longer needed?
I feel like the post goes from a fairly anthropomorphic approach of asking essentially—why bacteria failed to evolve into more complex forms. But from a non-anthropomorphic perspective, they failed nothing at all. They are highly resilient, persistent, widespread, adaptable, biologically successful in other terms, lifeforms. Rugged and simple—those designs tend to work. And to go back to everybody’s favourite topic—i.e. AI and the future that goes with it, or not—I would put their chances of being around in one thousand year well, well higher than those of homo sapiens—complex as it may be.
A person on Twitter reached out to me. I’m copying their response below:
Hello, I’m not on Lesswrong, but this is in response to your comment on “Why Are Bacteria So Simple?” I’m also not on Twitter much.
AlphaandOmega has it right, it’s random mutation/losses + evolutionary pressure. With plasmids loss is more common since if during division sometimes one cell ends up with no copies of the plasmid, and if the plasmid is costly then the one without the plasmid reproduces more quickly, and they can outcompete the bacteria with plasmids in hours.
(This is why many plasmids evolve partitioning systems or addiction modules to either make it more likely that both daughter cells have the plasmid, or to kill any daughters without plasmids. Plasmids are selfish, too!)
With resistance on the chromosome it gets a bit more complicated. New gene mutations (like mutating the target of the antibiotic) usually stick around, and the bacteria often evolves compensatory mutations to compensate for the reduced efficiency, though the mutation can still be “lost” if an unmutated bacteria still exists and can outcompete.
However, transposon and recombination can also be used to transfer resistance genes between cell chromosomes. Recombination and transposons often occur at specific sites due to having targeting sites for transposases, or higher AT content leading to weaker DNA bonds. But since these sites are hotspots of recombination, it’s also easier for DNA to be deleted from these regions as well.
There is a way to determine if a part of a genome is more “alien” than the rest, and that’s by looking at what’s called the GC content (percent of DNA that is G or C instead of A or T). If a region with otherwise similar GC content is interrupted by a region with a drastically different GC content, this can be evidence of a DNA insertion from a different species. It is unlikely that bacteria can detect this change and respond, but is interesting anyways.
--
The details are sound, but I’m not a big fan of the framing of the article; bacteria are far from simple, and undergo extremely complex regulation for survival under multiple environments (A single bacterium contains the energy metabolism contained in all eukaryotes, as well being able to use light to eat rocks, DMSO, and carbon monoxide, in addition to nitrogen fixation).
Bacteria also contain membrane-bound organelles (magnetosomes, for example), but membranes aren’t the only method of segregation, as you can have protein complexes or phase-separation boundaries to increase efficiency.
Bacteria also have “duplication and divergence”, in fact, probably moreso than eukaryotes. It has been shown that bacteria can duplicate copies of heavily-used genes, then delete them when they’re no longer required. But if the duplicates mutate before they’re deleted, they can diverge.
I’m not sure how accurate the section on modularity is; I know of bacterial systems which are controlled by three or four different regulators so that its levels can be precisely tuned for the environment it’s in. Not all bacterial genes are in operons, and even in operons there’s additional levels of regulation, including some not found in eukaryotes (like antitermination).
Bacteria also have their own high-level regulators, but since they generally lack multicellularity, it is often in response to specific states, like kinds of stress.
I will concede that bacteria have generally failed to evolve multicellularity and thus have not been able to achieve the specialization tradeoffs for things like body plans, though I think this issue is due to having a extra cell wall/lack of nucleus.
Why are bacteria so simple? They’re not. And given that we can only study the bacteria we know how to grow, I’d guess they’re even more complex than we thought.
I am no a biologist, but it seems to me that the most important difference between prokaryotes and eukaryotes is sexual reproduction rather than mitochondria (as I wrote about meanderingly). But neither article can resolve the issue, as my article ignores energy/mitochondria and yours ignores sex.
Still, it feels to me like this article is picking causes and effects kind of arbitrarily: “organism size” and “mitochondria” are taken to be a cause while “genome size” is taken to be an effect, but I don’t see you trying to justify the presence or direction of your arrows of causation.
I read Nick Lane’s The Vital Question and he argues that mitochondria came first and sex second. IIRC, he said that small-genome organisms (bacteria & archaea) do sporadic one-off gene exchange with their neighbors, and that process basically accomplishes the same thing that meiosis accomplishes in eukaryotes, i.e. enabling natural selection to act independently on every gene simultaneously. But sporadic one-off gene exchange is not a mechanism that can scale up to really massive numbers of genes. (I forget why not.) Hence, as genome size started expanding in early eukaryotes, they had to switch to meiosis (with crossing-over) at some point.
I have a vague recollection that he viewed meiosis as easy to evolve, with the justification that meiosis-related machinery can be found in some archaea (or something?). But very low confidence that I’m remembering correctly.
By contrast he definitely goes on and on about how you fundamentally can’t have a large genome without mitochondria [part of his argument is in the OP], and how the process of getting mitochondria (via an archaeon endocytosis-ing a bacterium) was an extremely difficult step evolutionarily.
Not all eukaryotes employ sexual reproduction. Also prokaryotes do have some mechanisms for DNA exchange as well, so copying errors are not their only chance for evolution either.
But I do agree that it’s probably no coincidence that the most complex life forms are sexually reproducing eukaryotes.
I’m pretty sure that I read (in Nick Lane’s The Vital Question) that all eukaryotes employ sexual reproduction at least sometimes. It’s true that they might reproduce asexually for a bunch of generations between sexual reproduction events. (It’s possible that other people disagree with Nick Lane on this, I dunno.)
All Eukaryotes Are Sexual, unless Proven Otherwise says maybe with enough hard work you can find some Malassezia fungi that are asexual, but also the title is pretty clear.
I see that someone strongly-disagreed with me on this. But are there any eukyrotes that cannot reproduce sexually (and are not very-recently-decended from sexual-reproducers) but still maintain size or complexity levels commonly associated with eukyrotes?
Indeed (as other commenters also pointed out) the ability to sexually reproduce seems to be much more prevalent than I originally thought when writing the above comment. (I thought that eukaryotes only capable of asexual reproduction were relatively common, but it seems that there may only be a very few special cases like that.)
I still disagree with you dismissing the importance of mitochondria though. (I don’t think the OP is saying that mitochondria alone are sufficient for larger genomes, but the argument for why they are at least necessary is convincing to me.)