AI’s impact on biology research: Part I, today
[12/29/23 edited to correct math error]
I’m a biology PhD, and have been working in tech for a number of years. I want to show why I believe that biological research is the most near term, high value application of machine learning. This has profound implications for human health, industrial development, and the fate of the world.
In this article I explain the current discoveries that machine learning has enabled in biology. In the next article I will consider what this implies will happen in the near term without major improvements in AI, along with my speculations about how our expectations that underlie our regulatory and business norms will fail. Finally, my last article will examine the longer term possibilities for machine learning and biology, including crazy but plausible sci-fi speculation.
TL;DR
Biology is complex, and the potential space of biological solutions to chemical, environmental, and other challenges is incredibly large. Biological research generates huge, well labeled datasets at low cost. This is a perfect fit with current machine learning approaches. Humans without computational assistance have very limited ability to understand biological systems enough to simulate, manipulate, and generate them. However, machine learning is giving us tools to do all of the above. This means things that have been constrained by human limits such as drug discovery or protein structure are suddenly unconstrained, turning a paucity of results into a superabundance in one step.
Biology and data
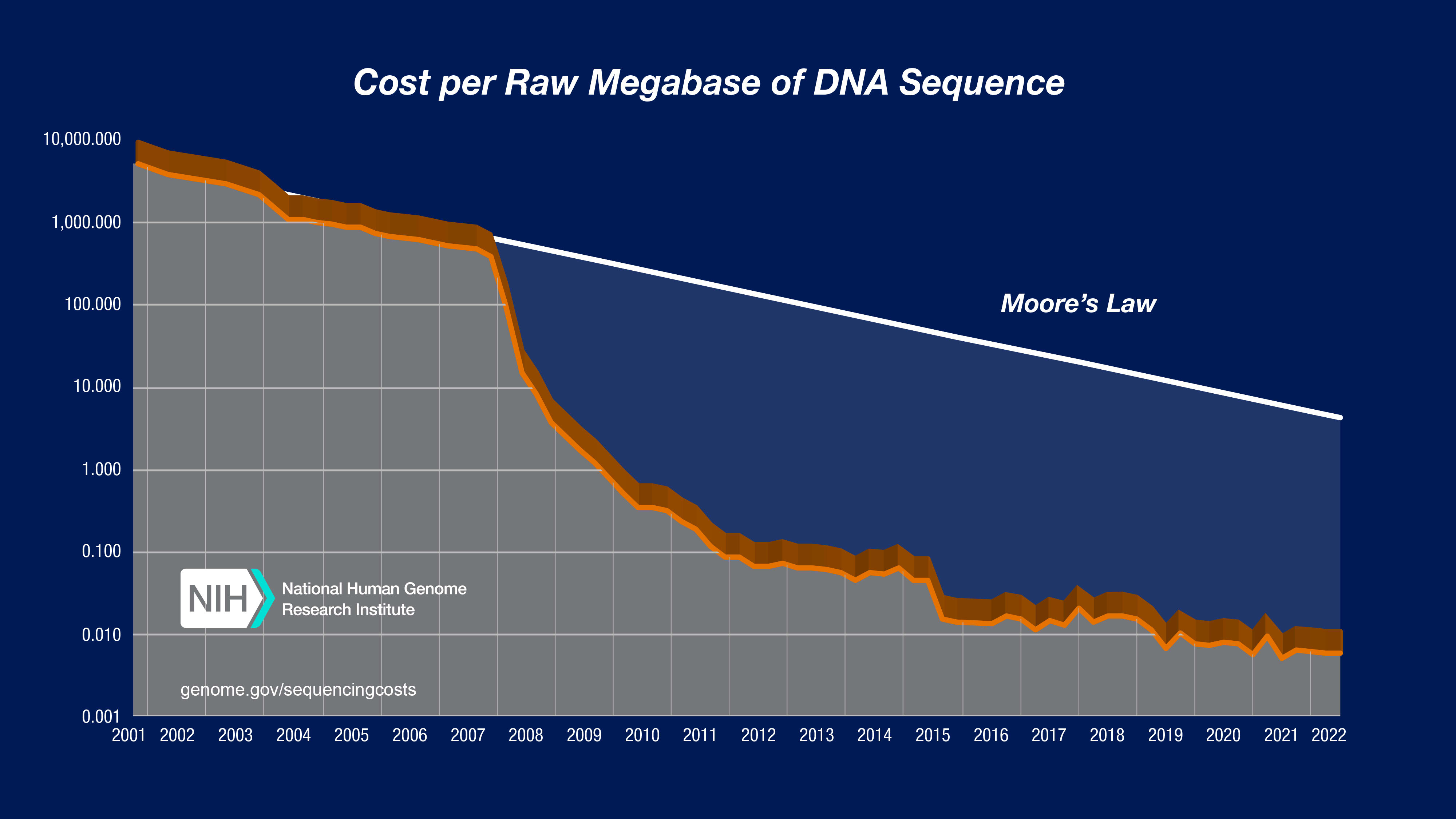
Biological research has been using technology to collect vast datasets since the bioinformatics revolution of the 1990′s. DNA sequencing costs have dropped by 5 orders of magnitude in 20 years ($100,000,000 dollars per human genome to $1000 dollars per genome)[1]. Microarrays allowed researchers to measure changes in mRNA expression in response to different experimental conditions across the entire genome of many species. High throughput cell sorting, robotic multi-well assays, proteomics chips, automated microscopy, and many more technologies generate petabytes of data.
As a result, biologists have been using computational tools to analyze and manipulate big datasets for over 30 years. Labs create, use, and share programs. Grad students are quick to adapt open source software, and lead researchers have been investing in powerful computational resources. There is a strong culture of adopting new technology, and this extends to machine learning.
Leading Machine Learning experts want to solve biology
Computer researchers have long been interested in applying computational resources to solve biological problems. Hedge fund billionaire David E. Shaw intentionally started a hedge fund so that he could fund computational biology research[2]. Demis Hassabis, Deepmind founder, is a PhD neuroscientist. Under his leadership Deepmind has made biological research a major priority, spinning off Isomorphic Labs[3] focused on drug discovery. The Chan Zuckerberg Institute is devoted to enabling computational research in biology and medicine to “cure, prevent, or manage all diseases by the end of this century”[4]. This shows that the highest level of machine learning research is being devoted to biological problems.
What have we discovered so far?
In 2020, Deepmind showed accuracy equal to the best physical methods of protein structure measurement at the CASP 14 protein folding prediction contest with their AlphaFold2 program.[5] This result “solved the protein folding problem”[6] for the large majority of proteins, showing that they could generate a high quality, biologically accurate 3D protein structure given the DNA sequence that encodes the protein. Deepmind then used AlphaFold2 to generate structures for all proteins known to humans, and contributed these structures to an open, free, public database. This increased the number of solved proteins available to researchers from ~180,000 to over 200,000,000[7]. Deepmind has continued to expand AlphaFold, adding mulit-protein complexes in 2022[8], and proteins and protein complexes interacting with DNA, RNA, and small molecules (such as drugs)[9].
The Baker lab at the University of Washington has used machine learning to create de novo proteins that bind to proteins in nature.[10] This allows biologists to create improved detection of proteins that may be rare in a sample. It also hints at therapeutic approaches that involve designer proteins or altered natural proteins as therapeutic agents.
The Collins Lab at the Broad Institute has used machine learning to design a new class of antibiotics.[11]
All of these results show that machine learning is solving long standing challenges in biology, and that these tools are being widely adopted. My next article will go into what we can expect in the near future, and some of the implications and likely disruptions that this will cause.
Small edit: $100,000,000 dollars per human genome to $1,000 dollars per genome is 5 orders of magnitude, not 6.
I’m looking forward to Part 2! (and Part 3?)
Thanks, I updated the article. Working on Part II now.
For those curious in the topic, here’s a related 80k podcast: https://80000hours.org/podcast/episodes/nita-farahany-neurotechnology/
Unfortunately, I believe we should anticipate the intersection of AI and biology to unlock novel dangers and dual-use things as well as progress on good things.
Indeed, there is a huge potential for dangerous applications with current technology. It is easier to design a molecule to poison a system than to heal a system.
Machine learning has great applications for biology. But beware that some of the media headlines are far too often focused on the intersection with Tech/Deep Learning, which aim mostly at solving a specific problem of biology. But it tells absolutely nothing about disease mechanism and does not answer why some combination of drugs can be synergistic or not in certain disease. It’s almost like engineers trying to find a use case to their fancy algorithm. And Tech VC pouring money into this because they understand the tech language but totally missed the point of patient survival.
Let’s say I use AI to generate an ideal small molecule as anti cancer drug. Nice, but what if the best way to make an impact in cancer is through cell therapy and not small molecule ? To use AI and ML for these applications is much harder from the scientific perspective, as we simply do not have enough quality data. I have some hope in Graph neural network, factor analysis and any basic ML, as long as everything is connected: Patient data (treatment+outcome) + genomics + In vitro/in vivo data
I totally agree. The techniques that have worked well so far are quite far from understanding cells, organs, organisms, or ecosystems. However, the incredible rate of progress at the small molecule to protein complex scale is already showing a strong impact, at least on getting a much better funnel before we begin in vivo testing.